Automatic differentiation with backward mode¶
NOTE
autodiff system in
pespaceandtiwaveis still under active development. If errors occur when running this notebook, try using the commit taggedautodiffmay solve the problem.
import numpy as np
from matplotlib import pyplot as plt
import bilby
import taichi as ti
ti.init(
arch=ti.cpu,
default_fp=ti.f64,
cpu_max_num_threads=1,
offline_cache=False,
debug=True,
unrolling_limit=0
)
[Taichi] version 1.7.4, llvm 15.0.4, commit b4b956fd, linux, python 3.10.19
[Taichi] Starting on arch=x64
[I 02/02/26 11:46:41.873 86535] [shell.py:_shell_pop_print@23] Graphical python shell detected, using wrapped sys.stdout
from pespace.detector.antenna import InterferometerAntenna, FDResponseModelMarset2018
from pespace.detector.tdi import TDIChannelData, FDMichelsonConstantEqualArm
from pespace.detector.orbit import KaplerianHeliocentric
from tiwave.waveforms import IMRPhenomXAS
tdi_gen = "2.0"
tdi_chan = ("A", "E", "T")
DAYJUL_SI = 86400.0
delta_time = 10
f_min = 1e-4
f_max = 0.5*(1/delta_time)
f_ref = f_min
t_start = 0.0
num_tsamples = 2 ** np.ceil(np.log2(30 * DAYJUL_SI/delta_time)) # two weeks
duration = num_tsamples * delta_time
before_tc = 0.8 * duration
after_tc = 0.2 * duration
tc = t_start + before_tc
print("sample num: ", num_tsamples)
print("duration: ", duration)
print("duration(day): ", duration/DAYJUL_SI)
print("tc: ", tc)
params = dict(
total_mass=3e6,
mass_ratio=0.6,
chi_1=0.75,
chi_2=0.62,
luminosity_distance=56000.0,
inclination=0.4,
reference_phase=1.3,
ecliptic_longitude=1.375,
ecliptic_latitude=-1.2108,
polarization=2.659,
coalescence_time=tc,
)
params = bilby.gw.conversion.generate_component_masses(params)
display(params)
params.pop('total_mass')
params.pop('mass_ratio')
sample num: 262144.0
duration: 2621440.0
duration(day): 30.340740740740742
tc: 2097152.0
{'total_mass': 3000000.0,
'mass_ratio': 0.6,
'chi_1': 0.75,
'chi_2': 0.62,
'luminosity_distance': 56000.0,
'inclination': 0.4,
'reference_phase': 1.3,
'ecliptic_longitude': 1.375,
'ecliptic_latitude': -1.2108,
'polarization': 2.659,
'coalescence_time': np.float64(2097152.0),
'mass_1': 1875000.0,
'mass_2': 1125000.0}
0.6
tdi_data = TDIChannelData()
tdi_data.set_fd_data_from_zero(
tdi_chan,
duration,
delta_time,
start_time=t_start,
minimum_frequency=f_min,
maximum_frequency=f_max,
)
tdi_data.set_fd_noise_power_density_from_model("LISA_SciRDv1", tdi_generation=tdi_gen)
orbit_model = KaplerianHeliocentric(2.5e9, 0.0, 0.0)
response_model = FDResponseModelMarset2018()
tdi_combination = FDMichelsonConstantEqualArm(generation=tdi_gen, orthogonal=True)
lisa = InterferometerAntenna(
name="lisa",
tdi_data=tdi_data,
orbit_model=orbit_model,
response_model=response_model,
tdi_combination=tdi_combination,
needs_grad=True,
)
waveform_tiw = IMRPhenomXAS(
tdi_data.frequency_samples,
f_ref,
needs_grad=True,
)
waveform_tiw.update_waveform(params)
lisa.update_detector_response(
waveform_tiw.waveform_container,
params["ecliptic_longitude"],
params["ecliptic_latitude"],
params["polarization"],
params["coalescence_time"],
)
/home/nrui/disk_ext/workspace/tiwave/tiwave/waveforms/base_waveform.py:74: UserWarning: check_parameters is disable, make sure all parameters passed in are valid.
warnings.warn(
# # abs
# plt.figure()
# plt.loglog(
# lisa.tdi_data.data_info.frequency_samples_array,
# np.abs(lisa.tdi_response_numpy["A"]),
# label="A",
# )
# plt.loglog(
# lisa.tdi_data.data_info.frequency_samples_array,
# np.abs(lisa.tdi_response_numpy["E"]),
# label="E",
# )
# plt.loglog(
# lisa.tdi_data.data_info.frequency_samples_array,
# np.abs(lisa.tdi_response_numpy["T"]),
# label="T",
# )
# plt.xlim(f_min, f_max)
# plt.legend()
# # real part
# plt.figure()
# plt.semilogx(
# lisa.tdi_data.data_info.frequency_samples_array,
# lisa.tdi_response_numpy["A"].real,
# label="A real",
# )
# plt.semilogx(
# lisa.tdi_data.data_info.frequency_samples_array,
# lisa.tdi_response_numpy["E"].real,
# label="E real",
# )
# plt.semilogx(
# lisa.tdi_data.data_info.frequency_samples_array,
# lisa.tdi_response_numpy["T"].real,
# label="T real",
# )
# plt.xlim(f_min, f_max)
# plt.legend()
# # imag part
# plt.figure()
# plt.semilogx(
# lisa.tdi_data.data_info.frequency_samples_array,
# lisa.tdi_response_numpy["A"].imag,
# label="A imag",
# )
# plt.semilogx(
# lisa.tdi_data.data_info.frequency_samples_array,
# lisa.tdi_response_numpy["E"].imag,
# label="E imag",
# )
# plt.semilogx(
# lisa.tdi_data.data_info.frequency_samples_array,
# lisa.tdi_response_numpy["T"].imag,
# label="T imag",
# )
# plt.xlim(f_min, f_max)
# plt.legend()
num = 10
iota_array = np.linspace(0.1, np.pi-0.1, num)
psds = lisa.tdi_data.fd_noise_power_density_numpy
delta_freq = lisa.tdi_data.data_info.delta_frequency
def get_rho_numpy(response, psds, df):
rho2 = 0.0
for chan in response.keys():
rho2 += np.vdot(response[chan], response[chan]/psds[chan]).real * 4.0 * df
return np.sqrt(rho2)
def get_d_rho_numpy(d_response, response, psds, df):
rho = get_rho_numpy(response, psds, df)
d_rho2 = 0.0
for chan in response.keys():
d_rho2 += np.vdot(d_response[chan], response[chan]/psds[chan]).real * df * 8
return d_rho2/(2*rho)
# symbolic differential
drho_diota_sd = np.zeros_like(iota_array)
@ti.kernel
def change_waveform_to_derivative(waveform_container: ti.template(), iota: float):
cos_iota = ti.cos(iota)
sin_iota = ti.sin(iota)
for i in waveform_container:
waveform_container[i].plus *= -(2*sin_iota*cos_iota)/(1+cos_iota*cos_iota)
waveform_container[i].cross *= -sin_iota/cos_iota
for idx in range(num):
params_sample = params.copy()
params_sample['inclination'] = iota_array[idx]
waveform_tiw.update_waveform(params_sample)
lisa.update_detector_response(waveform_tiw.waveform_container,
params_sample['ecliptic_longitude'],
params_sample['ecliptic_latitude'],
params_sample['polarization'],
params_sample['coalescence_time'],
)
response = lisa.tdi_response_numpy
change_waveform_to_derivative(waveform_tiw.waveform_container, params_sample['inclination'])
lisa.update_detector_response(waveform_tiw.waveform_container,
params_sample['ecliptic_longitude'],
params_sample['ecliptic_latitude'],
params_sample['polarization'],
params_sample['coalescence_time'],
)
d_response = lisa.tdi_response_numpy
d_rho = get_d_rho_numpy(d_response, response, psds, delta_freq)
drho_diota_sd[idx] = d_rho
drho_diota_sd
array([ -186.78885712, -744.5647822 , -1120.45507 , -1188.58411835,
-637.48134677, 642.05047074, 1191.29076756, 1122.63027232,
746.01518537, 187.15546285])
# numerical differential
drho_diota_nd_list = []
step_list = [1e-2, 1e-4, 1e-6, 1e-8, 1e-10, 1e-12]
for step in step_list:
drho_diota_nd = np.zeros_like(iota_array)
for idx in range(num):
params_left = params.copy()
params_left['inclination'] = iota_array[idx] - step
waveform_tiw.update_waveform(params_left)
lisa.update_detector_response(waveform_tiw.waveform_container,
params_left['ecliptic_longitude'],
params_left['ecliptic_latitude'],
params_left['polarization'],
params_left['coalescence_time'],
)
response_left = lisa.tdi_response_numpy
rho_left = get_rho_numpy(response_left, psds, delta_freq)
params_right = params.copy()
params_right['inclination'] = iota_array[idx] + step
waveform_tiw.update_waveform(params_right)
lisa.update_detector_response(waveform_tiw.waveform_container,
params_right['ecliptic_longitude'],
params_right['ecliptic_latitude'],
params_right['polarization'],
params_right['coalescence_time'],
)
response_right = lisa.tdi_response_numpy
rho_right = get_rho_numpy(response_right, psds, delta_freq)
drho_diota_nd[idx] = (rho_right - rho_left) / (2*step)
drho_diota_nd_list.append(drho_diota_nd)
drho_diota_nd_list
[array([ -186.78150276, -744.53597202, -1120.40875427, -1188.49544728,
-637.30592251, 641.87493609, 1191.20229988, 1122.58390764,
745.98631507, 187.14809241]),
array([ -186.78885639, -744.56477931, -1120.45506536, -1188.58410948,
-637.48132923, 642.05045318, 1191.29075872, 1122.63026769,
746.01518248, 187.15546211]),
array([ -186.78885726, -744.56478217, -1120.45507001, -1188.58411827,
-637.4813467 , 642.05047073, 1191.29076768, 1122.63027256,
746.01518565, 187.15546287]),
array([ -186.78886136, -744.56478387, -1120.45506739, -1188.58410474,
-637.48134949, 642.05046897, 1191.29076097, 1122.63027177,
746.01518918, 187.1554673 ]),
array([ -186.78747438, -744.56579569, -1120.45768219, -1188.58338283,
-637.4818895 , 642.05096351, 1191.29083487, 1122.63023766,
746.014166 , 187.15809347]),
array([ -186.78747438, -744.53510024, -1120.38378575, -1188.70957522,
-637.49894252, 642.04641603, 1191.3812159 , 1122.77120934,
746.2404028 , 187.12853489])]
# automatic differential
drho_diota_ad = np.zeros_like(iota_array)
rho2_ti = ti.field(dtype=ti.f64, shape=(), needs_grad=True)
rho_ti = ti.field(dtype=ti.f64, shape=(), needs_grad=True)
@ti.kernel
def get_rho_taichi():
rho_ti[None] = ti.sqrt(rho2_ti[None])
@ti.kernel
def get_rho2_taichi():
# reset rho2_ti outside of the kernel
for i in lisa.tdi_response:
inner_product = 0.0
for chan in ti.static(lisa.tdi_data.data_info.channels):
inner_product += (lisa.tdi_response[i][chan].norm_sqr()/lisa.tdi_data.fd_noise_power_density[i][chan] * 4.0 * delta_freq)
ti.atomic_add(rho2_ti[None], inner_product)
for idx in range(num):
params_sample = params.copy()
params_sample['inclination'] = iota_array[idx]
rho2_ti[None] = 0.0
rho_ti[None] = 0.0
with ti.ad.Tape(loss=rho_ti, clear_gradients=True):
waveform_tiw.update_waveform(params_sample)
lisa.update_detector_response(
waveform_tiw.waveform_container,
params_sample['ecliptic_longitude'],
params_sample['ecliptic_latitude'],
params_sample['polarization'],
params_sample['coalescence_time'],
)
get_rho2_taichi()
get_rho_taichi()
drho_diota_ad[idx] = waveform_tiw._params.iota.grad[None]
drho_diota_ad
array([ -186.78885712, -744.5647822 , -1120.45507 , -1188.58411835,
-637.48134677, 642.05047074, 1191.29076756, 1122.63027232,
746.01518537, 187.15546285])
import matplotlib
fig_width_pt = 3*246.0 # Get this from LaTeX using \showthe\columnwidth
inches_per_pt = 1.0/72.27 # Convert pt to inch
golden_mean = (np.sqrt(5)-1.0)/2.0 # Aesthetic ratio
fig_width = fig_width_pt*inches_per_pt # width in inches
fig_height = fig_width*golden_mean # height in inches
fig_size = [fig_width,fig_height]
plot_params = {'axes.labelsize': 24,
'font.family': 'serif',
'font.serif': 'Computer Modern',
'font.size': 24,
'legend.fontsize': 20,
'xtick.labelsize': 18,
'ytick.labelsize': 18,
'axes.grid' : False,
'text.usetex': True,
'savefig.dpi' : 100,
'lines.markersize' : 14,
'figure.figsize': fig_size}
matplotlib.rcParams.update(plot_params)
import os
os.environ['PATH'] = "/home/nrui/.local/texlive/2025/bin/x86_64-linux:" + os.environ['PATH']
color_list = ['tab:blue', 'tab:orange', 'tab:green', 'tab:red', 'tab:purple', 'tab:brown']
label_list = [r'ND(step=$10^{-2}$)', r'ND(step=$10^{-4}$)', r'ND(step=$10^{-6}$)', r'ND(step=$10^{-8}$)', r'ND(step=$10^{-10}$)', r'ND(step=$10^{-12}$)']
plt.figure(figsize=[fig_width*1.35,fig_height])
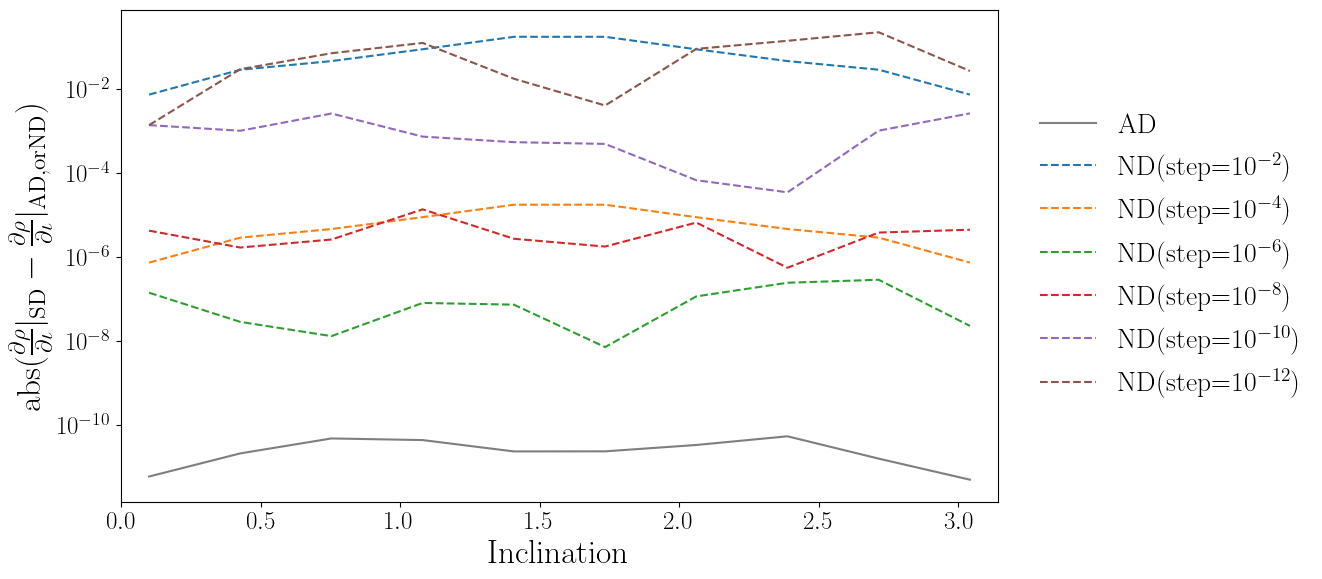
plt.semilogy(iota_array, np.abs(drho_diota_sd-drho_diota_ad), color='tab:gray', label='AD')
for idx, nd_data in enumerate(drho_diota_nd_list):
plt.semilogy(iota_array, np.abs(drho_diota_sd-nd_data), linestyle='dashed', color=color_list[idx], label=label_list[idx])
plt.xlabel("Inclination")
plt.ylabel(r'${\rm abs}(\frac{\partial \rho}{\partial \iota}|_{\rm SD} - \frac{\partial \rho}{\partial \iota}|_{\rm AD, or ND})$')
plt.xlim(0.0, np.pi)
plt.legend(
loc='center left',
bbox_to_anchor=(1.02, 0.5),
frameon=False
)
plt.tight_layout()
plt.savefig("numerical_err_diota_abs.pdf")
color_list = ['tab:blue', 'tab:orange', 'tab:green', 'tab:red', 'tab:purple', 'tab:brown']
label_list = [r'ND(step=$10^{-2}$)', r'ND(step=$10^{-4}$)', r'ND(step=$10^{-6}$)', r'ND(step=$10^{-8}$)', r'ND(step=$10^{-10}$)', r'ND(step=$10^{-12}$)']
plt.figure(figsize=[fig_width*1.35,fig_height])
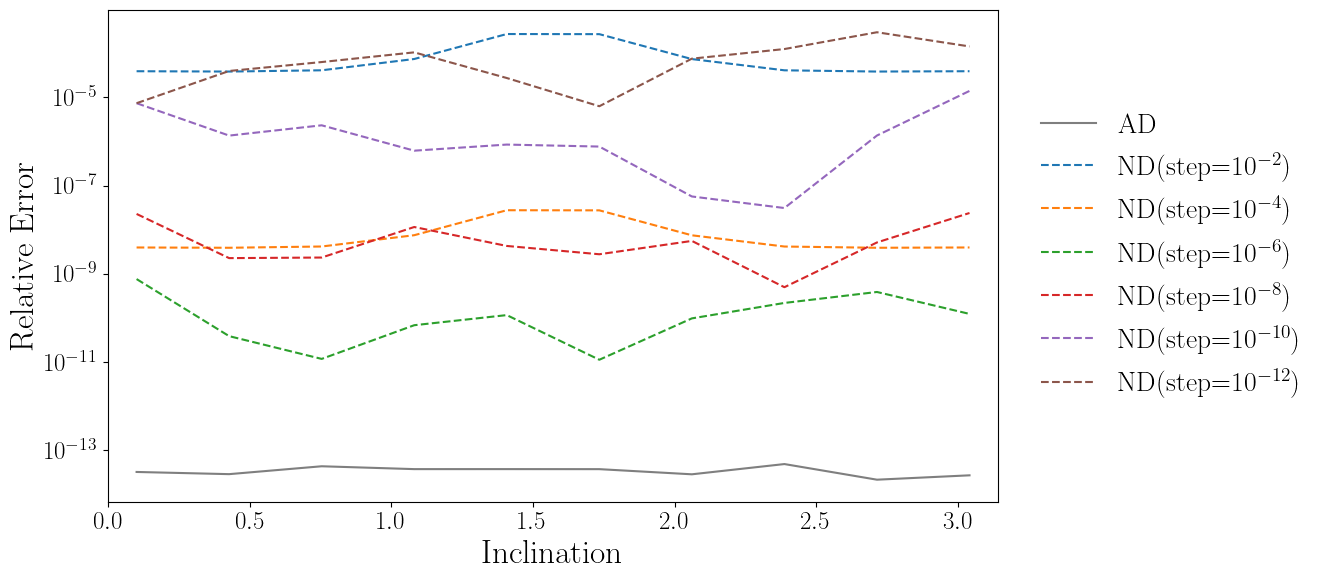
plt.semilogy(iota_array, np.abs((drho_diota_sd-drho_diota_ad)/drho_diota_sd), color='tab:gray', label='AD')
for idx, nd_data in enumerate(drho_diota_nd_list):
plt.semilogy(iota_array, np.abs((drho_diota_sd-nd_data)/drho_diota_sd), linestyle='dashed', color=color_list[idx], label=label_list[idx])
plt.xlabel("Inclination")
# plt.ylabel(r'${\rm abs}(\frac{\partial \rho}{\partial \iota}|_{\rm SD} - \frac{\partial \rho}{\partial \iota}|_{\rm AD, or ND})$')
# plt.ylabel(r'$\left| \frac{\frac{\partial \rho}{\partial \iota}|_{\rm SD} - \frac{\partial \rho}{\partial \iota}|_{\rm AD, or ND}}{\frac{\partial \rho}{\partial \iota}|_{\rm SD}}\right|$')
plt.ylabel("Relative Error")
plt.xlim(0.0, np.pi)
plt.legend(
loc='center left',
bbox_to_anchor=(1.02, 0.5),
frameon=False
)
plt.tight_layout()
plt.savefig("numerical_err_diota_rel.pdf")