Automatic differentiation with forward mode¶
NOTE
autodiff system in
pespaceandtiwaveis still under active development. If errors occur when running this notebook, try using the commit taggedautodiffmay solve the problem.
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
import bilby
import taichi as ti
ti.init(
arch=ti.cpu,
default_fp=ti.f64,
cpu_max_num_threads=1,
offline_cache=False,
# debug=True,
)
import os
os.environ['PATH'] = "/home/nrui/.local/texlive/2025/bin/x86_64-linux:" + os.environ['PATH']
[Taichi] version 1.7.4, llvm 15.0.4, commit b4b956fd, linux, python 3.10.19
[I 02/02/26 11:42:21.678 84923] [shell.py:_shell_pop_print@23] Graphical python shell detected, using wrapped sys.stdout
[Taichi] Starting on arch=x64
from pespace.detector.antenna import InterferometerAntenna, FDResponseModelMarset2018
from pespace.detector.tdi import TDIChannelData, FDMichelsonConstantEqualArm
from pespace.detector.orbit import available_orbit_models
from tiwave.waveforms import IMRPhenomXAS
tdi_gen = "2.0"
tdi_chan = ("A", "E", "T")
DAYJUL_SI = 86400.0
delta_time = 10
num_tsamples = 2 ** np.ceil(np.log2(7 * DAYJUL_SI / delta_time))
duration = num_tsamples * delta_time
print("sample num: ", num_tsamples)
print("duration: ", duration)
print("duration(day): ", duration/DAYJUL_SI)
f_min = 1e-4
f_ref = f_min
f_max = 0.5*(1/delta_time)
t_start = 0.0
before_tc = 0.8 * duration
after_tc = 0.2 * duration
tc = t_start + before_tc
params = dict(
total_mass=3e6,
mass_ratio=0.6,
chi_1=0.75,
chi_2=0.62,
luminosity_distance=56000.0,
inclination=0.4,
reference_phase=1.3,
ecliptic_longitude=1.375,
ecliptic_latitude=-1.2108,
polarization=2.659,
coalescence_time=tc,
)
params = bilby.gw.conversion.generate_mass_parameters(params)
print(params)
sample num: 65536.0
duration: 655360.0
duration(day): 7.5851851851851855
{'total_mass': 3000000.0, 'mass_ratio': 0.6, 'chi_1': 0.75, 'chi_2': 0.62, 'luminosity_distance': 56000.0, 'inclination': 0.4, 'reference_phase': 1.3, 'ecliptic_longitude': 1.375, 'ecliptic_latitude': -1.2108, 'polarization': 2.659, 'coalescence_time': np.float64(524288.0), 'mass_1': 1875000.0, 'mass_2': 1125000.0, 'chirp_mass': 1256226.717491785, 'symmetric_mass_ratio': np.float64(0.234375)}
tdi_data = TDIChannelData()
tdi_data.set_fd_data_from_zero(
tdi_chan,
duration,
delta_time,
start_time=t_start,
minimum_frequency=f_min,
maximum_frequency=f_max,
)
orbit_model = available_orbit_models['LISA_analytic']
response_model = FDResponseModelMarset2018()
tdi_combination = FDMichelsonConstantEqualArm(generation=tdi_gen, orthogonal=True)
lisa = InterferometerAntenna(
name="lisa",
tdi_data=tdi_data,
orbit_model=orbit_model,
response_model=response_model,
tdi_combination=tdi_combination,
needs_dual=True,
)
waveform_tiw = IMRPhenomXAS(
tdi_data.frequency_samples,
f_ref,
)
waveform_tiw.update_waveform(params)
lisa.update_detector_response(
waveform_tiw.waveform_container,
params['ecliptic_longitude'],
params["ecliptic_latitude"],
params["polarization"],
params["coalescence_time"],
)
/home/nrui/disk_ext/workspace/tiwave/tiwave/waveforms/base_waveform.py:74: UserWarning: check_parameters is disable, make sure all parameters passed in are valid.
warnings.warn(
# # abs
# plt.figure()
# plt.loglog(
# lisa.tdi_data.data_info.frequency_samples_array,
# np.abs(lisa.tdi_response_numpy["A"]),
# label="A",
# )
# plt.loglog(
# lisa.tdi_data.data_info.frequency_samples_array,
# np.abs(lisa.tdi_response_numpy["E"]),
# label="E",
# )
# plt.loglog(
# lisa.tdi_data.data_info.frequency_samples_array,
# np.abs(lisa.tdi_response_numpy["T"]),
# label="T",
# )
# plt.xlim(f_min, f_max)
# plt.legend()
# # real part
# plt.figure()
# plt.semilogx(
# lisa.tdi_data.data_info.frequency_samples_array,
# lisa.tdi_response_numpy["A"].real,
# label="A real",
# )
# plt.semilogx(
# lisa.tdi_data.data_info.frequency_samples_array,
# lisa.tdi_response_numpy["E"].real,
# label="E real",
# )
# plt.semilogx(
# lisa.tdi_data.data_info.frequency_samples_array,
# lisa.tdi_response_numpy["T"].real,
# label="T real",
# )
# plt.xlim(f_min, f_max)
# plt.legend()
# # imag part
# plt.figure()
# plt.semilogx(
# lisa.tdi_data.data_info.frequency_samples_array,
# lisa.tdi_response_numpy["A"].imag,
# label="A imag",
# )
# plt.semilogx(
# lisa.tdi_data.data_info.frequency_samples_array,
# lisa.tdi_response_numpy["E"].imag,
# label="E imag",
# )
# plt.semilogx(
# lisa.tdi_data.data_info.frequency_samples_array,
# lisa.tdi_response_numpy["T"].imag,
# label="T imag",
# )
# plt.xlim(f_min, f_max)
# plt.legend()
field_names = [
'link12_re',
'link12_im',
'link21_re',
'link21_im',
'link23_re',
'link23_im',
'link32_re',
'link32_im',
'link31_re',
'link31_im',
'link13_re',
'link13_im',
]
loss = {key: ti.field(dtype=ti.f64, shape=tdi_data.frequency_samples.shape, needs_dual=True) for key in field_names}
# print(loss)
# print(loss['link12_re'])
# @ti.kernel
# def test():
# print(ti.get_addr(loss['link12_re'], 0))
# print(ti.get_addr(loss['link12_im'], 0))
# print(ti.get_addr(loss['link21_re'], 0))
# print(ti.get_addr(loss['link21_im'], 0))
# print(ti.get_addr(loss['link23_re'], 0))
# print(ti.get_addr(loss['link23_im'], 0))
# print(ti.get_addr(loss['link32_re'], 0))
# print(ti.get_addr(loss['link32_im'], 0))
# print(ti.get_addr(loss['link31_re'], 0))
# print(ti.get_addr(loss['link31_im'], 0))
# print(ti.get_addr(loss['link13_re'], 0))
# print(ti.get_addr(loss['link13_im'], 0))
# test()
@ti.kernel
def get_loss():
for i in lisa.single_link_response:
loss['link12_re'][i] = lisa.single_link_response[i]['link12'][0]
loss['link12_im'][i] = lisa.single_link_response[i]['link12'][1]
loss['link21_re'][i] = lisa.single_link_response[i]['link21'][0]
loss['link21_im'][i] = lisa.single_link_response[i]['link21'][1]
loss['link23_re'][i] = lisa.single_link_response[i]['link23'][0]
loss['link23_im'][i] = lisa.single_link_response[i]['link23'][1]
loss['link32_re'][i] = lisa.single_link_response[i]['link32'][0]
loss['link32_im'][i] = lisa.single_link_response[i]['link32'][1]
loss['link31_re'][i] = lisa.single_link_response[i]['link31'][0]
loss['link31_im'][i] = lisa.single_link_response[i]['link31'][1]
loss['link13_re'][i] = lisa.single_link_response[i]['link13'][0]
loss['link13_im'][i] = lisa.single_link_response[i]['link13'][1]
get_loss()
# print(loss['link12_re'])
# print(loss['link12_im'])
# print(loss['link21_re'])
# print(loss['link21_im'])
# print(loss['link23_re'])
# print(loss['link23_im'])
# print(loss['link32_re'])
# print(loss['link32_im'])
# print(loss['link31_re'])
# print(loss['link31_im'])
# print(loss['link13_re'])
# print(loss['link13_im'])
loss_list = list(loss.values())
# print(loss_list)
# @ti.kernel
# def test():
# print(ti.get_addr(loss_list[0], 0)- ti.get_addr(loss['link12_re'], 0))
# print(ti.get_addr(loss_list[1], 0)- ti.get_addr(loss['link12_im'], 0))
# print(ti.get_addr(loss_list[2], 0)- ti.get_addr(loss['link21_re'], 0))
# print(ti.get_addr(loss_list[3], 0)- ti.get_addr(loss['link21_im'], 0))
# print(ti.get_addr(loss_list[4], 0)- ti.get_addr(loss['link23_re'], 0))
# print(ti.get_addr(loss_list[5], 0)- ti.get_addr(loss['link23_im'], 0))
# print(ti.get_addr(loss_list[6], 0)- ti.get_addr(loss['link32_re'], 0))
# print(ti.get_addr(loss_list[7], 0)- ti.get_addr(loss['link32_im'], 0))
# print(ti.get_addr(loss_list[8], 0)- ti.get_addr(loss['link31_re'], 0))
# print(ti.get_addr(loss_list[9], 0)- ti.get_addr(loss['link31_im'], 0))
# print(ti.get_addr(loss_list[10], 0)- ti.get_addr(loss['link13_re'], 0))
# print(ti.get_addr(loss_list[11], 0)- ti.get_addr(loss['link13_im'], 0))
# test()
with ti.ad.FwdMode(
loss=loss_list,
param=lisa.params.lam,
clear_gradients=True,
):
lisa.update_input_params(
params['ecliptic_longitude'],
params['ecliptic_latitude'],
params['polarization'],
params['coalescence_time'],
)
lisa.response_model._update_geometry_terms()
lisa.response_model._loop_frequencies(waveform_tiw.waveform_container)
get_loss()
dlinks_dlam = {key: loss[key].dual.to_numpy() for key in field_names}
display(dlinks_dlam)
{'link12_re': array([-8.49490177e-18, 8.97838709e-18, 1.04649853e-18, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00], shape=(32703,)),
'link12_im': array([-4.07607627e-18, -2.12022433e-18, 8.98819846e-18, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00], shape=(32703,)),
'link21_re': array([-8.49493389e-18, 8.97841965e-18, 1.04650619e-18, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00], shape=(32703,)),
'link21_im': array([-4.07608753e-18, -2.12023581e-18, 8.98823266e-18, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00], shape=(32703,)),
'link23_re': array([ 1.22286651e-17, -1.84576347e-17, 3.74647530e-18, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00], shape=(32703,)),
'link23_im': array([ 1.34622121e-17, -2.19153240e-18, -1.85876021e-17, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00], shape=(32703,)),
'link32_re': array([ 1.22286205e-17, -1.84575651e-17, 3.74645995e-18, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00], shape=(32703,)),
'link32_im': array([ 1.34621617e-17, -2.19152322e-18, -1.85875307e-17, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00], shape=(32703,)),
'link31_re': array([-9.89016235e-18, -1.27974159e-17, 3.01154492e-17, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00], shape=(32703,)),
'link31_im': array([ 3.09011604e-17, -2.94919898e-17, -1.03816627e-17, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00], shape=(32703,)),
'link13_re': array([-9.89017769e-18, -1.27973963e-17, 3.01154463e-17, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00], shape=(32703,)),
'link13_im': array([ 3.09011497e-17, -2.94919902e-17, -1.03816423e-17, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00], shape=(32703,))}
with ti.ad.FwdMode(
loss=loss_list,
param=lisa.params.beta,
clear_gradients=True,
):
lisa.update_input_params(
params['ecliptic_longitude'],
params['ecliptic_latitude'],
params['polarization'],
params['coalescence_time'],
)
lisa.response_model._update_geometry_terms()
lisa.response_model._loop_frequencies(waveform_tiw.waveform_container)
get_loss()
dlinks_dbeta = {key: loss[key].dual.to_numpy() for key in field_names}
display(dlinks_dbeta)
{'link12_re': array([ 1.74292183e-17, -1.54531410e-17, -6.72871199e-18, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00], shape=(32703,)),
'link12_im': array([ 3.49193585e-18, 8.39199199e-18, -1.60499126e-17, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00], shape=(32703,)),
'link21_re': array([ 1.74292849e-17, -1.54531990e-17, -6.72874374e-18, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00], shape=(32703,)),
'link21_im': array([ 3.49194400e-18, 8.39202916e-18, -1.60499757e-17, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00], shape=(32703,)),
'link23_re': array([-1.27180069e-17, 1.57120126e-18, 1.76435171e-17, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00], shape=(32703,)),
'link23_im': array([ 1.23954677e-17, -1.77758035e-17, 3.09839795e-18, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00], shape=(32703,)),
'link32_re': array([-1.27179742e-17, 1.57119864e-18, 1.76434702e-17, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00], shape=(32703,)),
'link32_im': array([ 1.23954336e-17, -1.77757562e-17, 3.09839123e-18, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00], shape=(32703,)),
'link31_re': array([ 2.05466738e-18, 7.04665982e-20, -4.09424760e-18, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00], shape=(32703,)),
'link31_im': array([-2.83442664e-18, 3.81499045e-18, -4.89672838e-19, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00], shape=(32703,)),
'link13_re': array([ 2.05473360e-18, 7.04009315e-20, -4.09427170e-18, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00], shape=(32703,)),
'link13_im': array([-2.83440563e-18, 3.81501909e-18, -4.89742538e-19, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00], shape=(32703,))}
with ti.ad.FwdMode(
loss=loss_list,
param=lisa.params.psi,
clear_gradients=True,
):
lisa.update_input_params(
params['ecliptic_longitude'],
params['ecliptic_latitude'],
params['polarization'],
params['coalescence_time'],
)
lisa.response_model._update_geometry_terms()
lisa.response_model._loop_frequencies(waveform_tiw.waveform_container)
get_loss()
dlinks_dpsi = {key: loss[key].dual.to_numpy() for key in field_names}
display(dlinks_dpsi)
{'link12_re': array([-1.10271623e-17, 7.03944628e-18, 7.36166411e-18, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00], shape=(32703,)),
'link12_im': array([ 2.00917615e-18, -8.34250435e-18, 7.69148073e-18, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00], shape=(32703,)),
'link21_re': array([-1.10272051e-17, 7.03947456e-18, 7.36169471e-18, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00], shape=(32703,)),
'link21_im': array([ 2.00918395e-18, -8.34253787e-18, 7.69151271e-18, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00], shape=(32703,)),
'link23_re': array([ 8.57510576e-18, -1.37165243e-17, 3.38739088e-18, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00], shape=(32703,)),
'link23_im': array([ 1.03804313e-17, -2.23595995e-18, -1.39027787e-17, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00], shape=(32703,)),
'link32_re': array([ 8.57507399e-18, -1.37164724e-17, 3.38737782e-18, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00], shape=(32703,)),
'link32_im': array([ 1.03803928e-17, -2.23595150e-18, -1.39027251e-17, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00], shape=(32703,)),
'link31_re': array([-1.09757867e-17, -1.50925982e-17, 3.45831495e-17, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00], shape=(32703,)),
'link31_im': array([ 3.53884296e-17, -3.36857897e-17, -1.24831449e-17, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00], shape=(32703,)),
'link13_re': array([-1.09757848e-17, -1.50925946e-17, 3.45831391e-17, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00], shape=(32703,)),
'link13_im': array([ 3.53884234e-17, -3.36857817e-17, -1.24831411e-17, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00], shape=(32703,))}
with ti.ad.FwdMode(
loss=loss_list,
param=lisa.params.tc,
clear_gradients=True,
):
lisa.update_input_params(
params['ecliptic_longitude'],
params['ecliptic_latitude'],
params['polarization'],
params['coalescence_time'],
)
lisa.response_model._update_geometry_terms()
lisa.response_model._loop_frequencies(waveform_tiw.waveform_container)
get_loss()
dlinks_dtc = {key: loss[key].dual.to_numpy() for key in field_names}
display(dlinks_dtc)
{'link12_re': array([ 3.47582264e-21, -2.24881505e-21, -2.39497177e-21, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00], shape=(32703,)),
'link12_im': array([-6.39347795e-22, 2.67370770e-21, -2.49479841e-21, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00], shape=(32703,)),
'link21_re': array([ 3.47583612e-21, -2.24882409e-21, -2.39498173e-21, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00], shape=(32703,)),
'link21_im': array([-6.39350273e-22, 2.67371844e-21, -2.49480878e-21, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00], shape=(32703,)),
'link23_re': array([-2.70230965e-21, 4.40593580e-21, -1.12289457e-21, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00], shape=(32703,)),
'link23_im': array([-3.29786288e-21, 7.36201860e-22, 4.53049074e-21, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00], shape=(32703,)),
'link32_re': array([-2.70229964e-21, 4.40591915e-21, -1.12289024e-21, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00], shape=(32703,)),
'link32_im': array([-3.29785066e-21, 7.36199080e-22, 4.53047327e-21, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00], shape=(32703,)),
'link31_re': array([ 3.46233528e-21, 4.88221890e-21, -1.13029852e-20, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00], shape=(32703,)),
'link31_im': array([-1.12380909e-20, 1.08451503e-20, 4.10133955e-21, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00], shape=(32703,)),
'link13_re': array([ 3.46233467e-21, 4.88221775e-21, -1.13029818e-20, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00], shape=(32703,)),
'link13_im': array([-1.12380889e-20, 1.08451477e-20, 4.10133833e-21, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00], shape=(32703,))}
links_name = [
'link12', 'link21', 'link23', 'link32', 'link31', 'link13'
]
stacked_dlinks_dlam = {
key: np.stack((dlinks_dlam[f'{key}_re'], dlinks_dlam[f'{key}_im']), axis=-1) for key in links_name
}
stacked_dlinks_dbeta = {
key: np.stack((dlinks_dbeta[f'{key}_re'], dlinks_dbeta[f'{key}_im']), axis=-1) for key in links_name
}
stacked_dlinks_dpsi = {
key: np.stack((dlinks_dpsi[f'{key}_re'], dlinks_dpsi[f'{key}_im']), axis=-1) for key in links_name
}
stacked_dlinks_dtc = {
key: np.stack((dlinks_dtc[f'{key}_re'], dlinks_dtc[f'{key}_im']), axis=-1) for key in links_name
}
lisa.single_link_response.from_numpy(stacked_dlinks_dlam)
lisa.tdi_combination.update_tdi_response()
dtdi_dlam = lisa.tdi_response_numpy
lisa.single_link_response.from_numpy(stacked_dlinks_dbeta)
lisa.tdi_combination.update_tdi_response()
dtdi_dbeta = lisa.tdi_response_numpy
lisa.single_link_response.from_numpy(stacked_dlinks_dpsi)
lisa.tdi_combination.update_tdi_response()
dtdi_dpsi = lisa.tdi_response_numpy
lisa.single_link_response.from_numpy(stacked_dlinks_dtc)
lisa.tdi_combination.update_tdi_response()
dtdi_dtc = lisa.tdi_response_numpy
display(dtdi_dlam)
display(dtdi_dbeta)
display(dtdi_dpsi)
display(dtdi_dtc)
{'A': array([-7.10068803e-21+1.66449185e-20j, -5.56315182e-21-1.76462038e-20j,
1.84627577e-20-4.08498861e-21j, ...,
0.00000000e+00+0.00000000e+00j, 0.00000000e+00+0.00000000e+00j,
0.00000000e+00+0.00000000e+00j], shape=(32703,)),
'E': array([-1.14813652e-20-9.35828789e-21j, 1.54238995e-20-2.49107373e-22j,
-1.25955113e-21+1.59975704e-20j, ...,
0.00000000e+00+0.00000000e+00j, 0.00000000e+00+0.00000000e+00j,
0.00000000e+00+0.00000000e+00j], shape=(32703,)),
'T': array([-4.79289871e-29+6.33746301e-29j, -3.54035743e-30-8.63423893e-29j,
9.36272322e-29+6.03547618e-30j, ...,
0.00000000e+00+0.00000000e+00j, 0.00000000e+00+0.00000000e+00j,
0.00000000e+00+0.00000000e+00j], shape=(32703,))}
{'A': array([-3.14946505e-22-6.78548779e-21j, 4.65475384e-21+5.43661196e-21j,
-6.31036552e-21+4.12401224e-21j, ...,
0.00000000e+00+0.00000000e+00j, 0.00000000e+00+0.00000000e+00j,
0.00000000e+00+0.00000000e+00j], shape=(32703,)),
'E': array([ 1.63555490e-20-5.16059499e-21j, -9.29442891e-21+1.48896665e-20j,
-1.43237935e-20-1.08193201e-20j, ...,
0.00000000e+00+0.00000000e+00j, 0.00000000e+00+0.00000000e+00j,
0.00000000e+00+0.00000000e+00j], shape=(32703,)),
'T': array([-5.42725447e-30-1.12252591e-28j, 8.19435763e-29+8.81232541e-29j,
-1.02078427e-28+7.82089273e-29j, ...,
0.00000000e+00+0.00000000e+00j, 0.00000000e+00+0.00000000e+00j,
0.00000000e+00+0.00000000e+00j], shape=(32703,))}
{'A': array([-5.80158628e-21+1.85021096e-20j, -7.97536192e-21-1.82895613e-20j,
1.94104900e-20-6.64176125e-21j, ...,
0.00000000e+00+0.00000000e+00j, 0.00000000e+00+0.00000000e+00j,
0.00000000e+00+0.00000000e+00j], shape=(32703,)),
'E': array([-1.07773430e-20-4.36898635e-21j, 1.16035954e-20-3.65158227e-21j,
2.53928108e-21+1.24604035e-20j, ...,
0.00000000e+00+0.00000000e+00j, 0.00000000e+00+0.00000000e+00j,
0.00000000e+00+0.00000000e+00j], shape=(32703,)),
'T': array([-2.40736848e-29+4.97428060e-29j, -1.32239375e-29-5.91944225e-29j,
6.62004769e-29-6.53900078e-30j, ...,
0.00000000e+00+0.00000000e+00j, 0.00000000e+00+0.00000000e+00j,
0.00000000e+00+0.00000000e+00j], shape=(32703,))}
{'A': array([ 1.82941921e-24-5.87450826e-24j, 2.58026074e-24+5.88673999e-24j,
-6.34222092e-24+2.18316638e-24j, ...,
0.00000000e+00+0.00000000e+00j, 0.00000000e+00+0.00000000e+00j,
0.00000000e+00+0.00000000e+00j], shape=(32703,)),
'E': array([ 3.39694478e-24+1.38796247e-24j, -3.72054353e-24+1.15931654e-24j,
-8.14018032e-25-4.05398969e-24j, ...,
0.00000000e+00+0.00000000e+00j, 0.00000000e+00+0.00000000e+00j,
0.00000000e+00+0.00000000e+00j], shape=(32703,)),
'T': array([ 7.64444793e-33-1.56719091e-32j, 4.17530253e-33+1.89786331e-32j,
-2.15377567e-32+2.05440990e-33j, ...,
0.00000000e+00+0.00000000e+00j, 0.00000000e+00+0.00000000e+00j,
0.00000000e+00+0.00000000e+00j], shape=(32703,))}
tdi_data.set_fd_noise_power_density_from_model("LISA_SciRDv1", tdi_generation=tdi_gen)
psds = lisa.tdi_data.fd_noise_power_density_numpy
def get_fisher_matrix(derivative_data, psd_data, df):
size = len(derivative_data)
fm = np.zeros((size, size))
for i in range(size):
for j in range(size):
inner_prod = np.vdot(derivative_data[i], derivative_data[j]/psd_data).real
inner_prod *= 4*df
fm[i, j] = inner_prod
return fm
fm_A = get_fisher_matrix(
[dtdi_dlam['A'], dtdi_dbeta['A'], dtdi_dpsi['A'], dtdi_dtc['A']],
psds['A'],
tdi_data.data_info.delta_frequency,
)
fm_E = get_fisher_matrix(
[dtdi_dlam['E'], dtdi_dbeta['E'], dtdi_dpsi['E'], dtdi_dtc['E']],
psds['E'],
tdi_data.data_info.delta_frequency,
)
fm_T = get_fisher_matrix(
[dtdi_dlam['T'], dtdi_dbeta['T'], dtdi_dpsi['T'], dtdi_dtc['T']],
psds['T'],
tdi_data.data_info.delta_frequency,
)
fm_comb_AET = fm_A + fm_E + fm_T
print(fm_comb_AET)
cov_AET = np.linalg.inv(fm_comb_AET)
print(cov_AET)
[[ 2.26317548e+07 1.01433958e+07 -1.08414299e+07 1.87855891e+05]
[ 1.01433958e+07 6.30512570e+06 -6.48278949e+06 9.30059279e+04]
[-1.08414299e+07 -6.48278949e+06 7.71973533e+06 -1.05111160e+05]
[ 1.87855891e+05 9.30059279e+04 -1.05111160e+05 1.66041851e+03]]
[[ 3.49566764e-06 -9.50811498e-07 -3.97684000e-06 -5.93983332e-04]
[-9.50811498e-07 1.56888845e-06 1.81335752e-06 1.34486412e-04]
[-3.97684000e-06 1.81335752e-06 5.87109893e-06 7.20021991e-04]
[-5.93983332e-04 1.34486412e-04 7.20021991e-04 1.05851374e-01]]
fm_comb_AE = fm_A + fm_E
print(fm_comb_AE)
cov_AE = np.linalg.inv(fm_comb_AE)
print(cov_AE)
[[ 2.26313505e+07 1.01431890e+07 -1.08412685e+07 1.87852599e+05]
[ 1.01431890e+07 6.30500845e+06 -6.48270061e+06 9.30042086e+04]
[-1.08412685e+07 -6.48270061e+06 7.71966395e+06 -1.05109804e+05]
[ 1.87852599e+05 9.30042086e+04 -1.05109804e+05 1.66039146e+03]]
[[ 3.49569527e-06 -9.50840551e-07 -3.97689180e-06 -5.93988594e-04]
[-9.50840551e-07 1.56892288e-06 1.81341322e-06 1.34491746e-04]
[-3.97689180e-06 1.81341322e-06 5.87120082e-06 7.20032112e-04]
[-5.93988594e-04 1.34491746e-04 7.20032112e-04 1.05852410e-01]]
samples = np.random.multivariate_normal(
[params['ecliptic_longitude'],
params['ecliptic_latitude'],
params['polarization'],
params['coalescence_time']],
cov_AET,
size=10000)
samples_pd = pd.DataFrame(
dict(ecliptic_longitude=samples[:,0],
ecliptic_latitude=samples[:,1],
polarization=samples[:,2],
coalescence_time=samples[:,3],
)
)
import matplotlib
params = {
'font.family': 'serif',
'font.serif': 'Computer Modern',
'legend.fontsize': 28,
'text.usetex': True,
}
matplotlib.rcParams.update(params)
fisher_res = bilby.core.result.Result(label='fisher_samples', search_parameter_keys=list(samples_pd.keys()), posterior=samples_pd)
# the posterior data can be found in https://doi.org/10.5281/zenodo.18339164
pe_res = bilby.core.result.read_in_result("intro-pespace/autodiff/pe_resp_params/outdir_extrinsic_params_LISA_noiseless/extrinsic_params_LISA_noiseless_result.json")
latex_labels = dict(
ecliptic_longitude=r"$\lambda$",
ecliptic_latitude=r"$\beta$",
polarization=r"$\psi$",
coalescence_time=r"$t_c$",
)
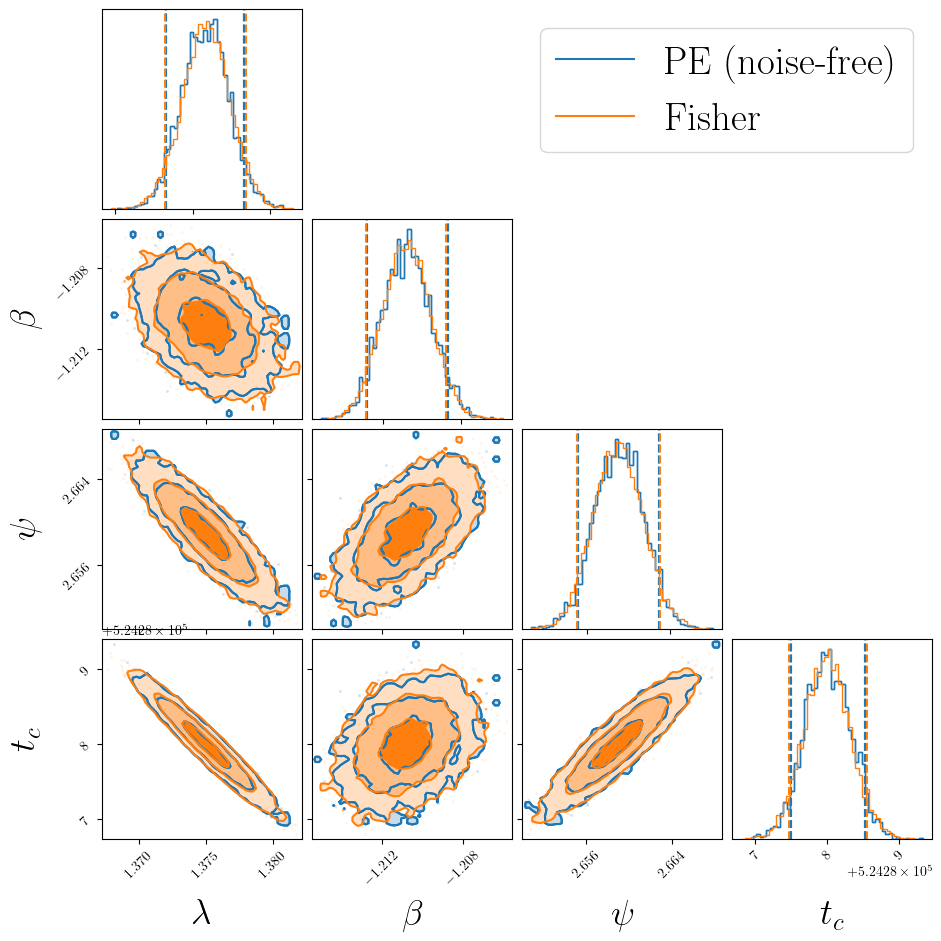
fig = bilby.core.result.plot_multiple(
[pe_res, fisher_res],
filename="corner_fisher.pdf",
labels=["PE (noise-free)", "Fisher"],
titles=False,
quantiles=[0.05, 0.95],
corner_labels=[latex_labels[key] for key in fisher_res.search_parameter_keys],
label_kwargs={'fontsize': 26}
)
params = {
'axes.labelsize': 18,
'font.family': 'serif',
'font.serif': 'Computer Modern',
'xtick.labelsize': 18,
'ytick.labelsize': 18,
'text.usetex': True,
'text.latex.preamble': r'\usepackage{amsmath}',
}
matplotlib.rcParams.update(params)
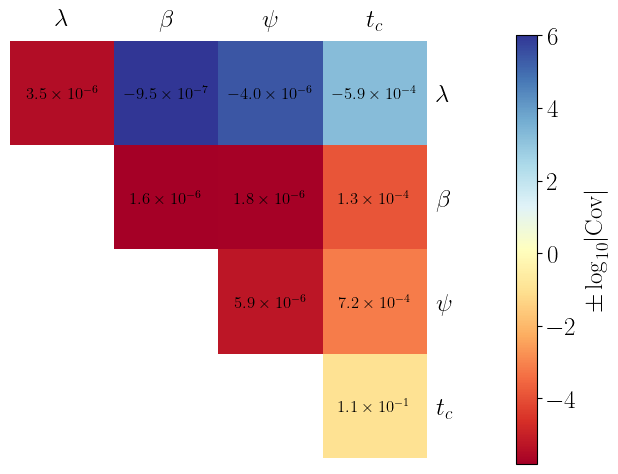
fig, ax = plt.subplots()
mask = np.tril(np.ones_like(cov_AET, dtype=bool), k=-1)
cov_masked = np.ma.array(cov_AET, mask=mask)
log10_cov_masked = np.sign(cov_masked) * np.log10(np.abs(cov_masked))
im = ax.imshow(log10_cov_masked, cmap="RdYlBu")
labels = [r'$\lambda$', r'$\beta$', r'$\psi$', r'$t_c$']
ax.set_xticks(range(len(labels)))
ax.set_yticks(range(len(labels)))
ax.set_xticklabels(labels)
ax.set_yticklabels(labels)
for spine in ax.spines.values():
spine.set_visible(False)
ax.xaxis.set_label_position('top')
ax.yaxis.set_label_position('right')
ax.xaxis.tick_top()
ax.yaxis.tick_right()
ax.tick_params(
top=False,
right=False,
bottom=False,
left=False
)
cbar = plt.colorbar(im, cmap="RdYlBu", pad=0.15)
cbar.set_label(r"$\pm \log_{10} \lvert \mathrm{Cov} \rvert$")
for i in range(4):
for j in range(i, 4):
val = cov_AET[i, j]
mantissa, exp = f"{val:.1e}".split('e')
exp = int(exp)
text = f"${mantissa}\\times10^{{{exp}}}$"
ax.text(
j, i,
text,
ha='center',
va='center',
fontsize=12,
color='black'
)
plt.tight_layout()
fig.savefig("cov.pdf")
cov_AET
array([[ 3.49566764e-06, -9.50811498e-07, -3.97684000e-06,
-5.93983332e-04],
[-9.50811498e-07, 1.56888845e-06, 1.81335752e-06,
1.34486412e-04],
[-3.97684000e-06, 1.81335752e-06, 5.87109893e-06,
7.20021991e-04],
[-5.93983332e-04, 1.34486412e-04, 7.20021991e-04,
1.05851374e-01]])